GENOMIC SELECTION AND INTROGRESSION SIGNATURES RESULTING FROM ADAPTATION TO HIGH SALINITY: A CASE STUDY ON AN INDONESIAN FARMED TILAPIA (SUKAMANDI) STRAIN
Background
Tilapia is currently the most important fish in aquaculture in the tropics and subtropics. Originally a freshwater species, it is cultured in a wide range of conditions. Among the most challenging is when high salinity is involved, e.g. culturing in estuaries or high salinity ponds in polyculture with shrimp. Optimized tilapia strains generally do not tolerate high salinity well. Although the physiological characteristics for osmoregulation are reasonably well understood, it is less clear how selection results in salinity tolerance. Here we investigate one such strain that was bred to perform well in brackish water. Specifically, we infer signatures of selections in the genome. In addition, since the salinity tolerance in the strain was hypothesized to be derived from another species, we also inferred signatures of introgression between species.
Materials and methods
Indonesian farmed tilapia (Sukamandi ) strain was derived from the BRPI research institute (Sukamandi , Java), and has been selected for growth for four generations under salinity levels varying from 15 to 58 ppt. Fin clips from a total of 20 fish were sampled from the nucleus population in 2019. In order to understand the genomic architecture of Sukamandi strain, whole genome sequencing data of blue tilapia were downloaded from the Sequence Read Archive (NCBI bio-project number PRJNA358089 and PRJEB23203). Nile tilapia, originating from the Genetically Improved Farmed Tilapia (GIFT) population, were collected by the Genomar company . We performed a joint-calling strategy for genetic variants, genetic differentiation and diversity analysis, selection signatures and Identity by descent detection.
Results
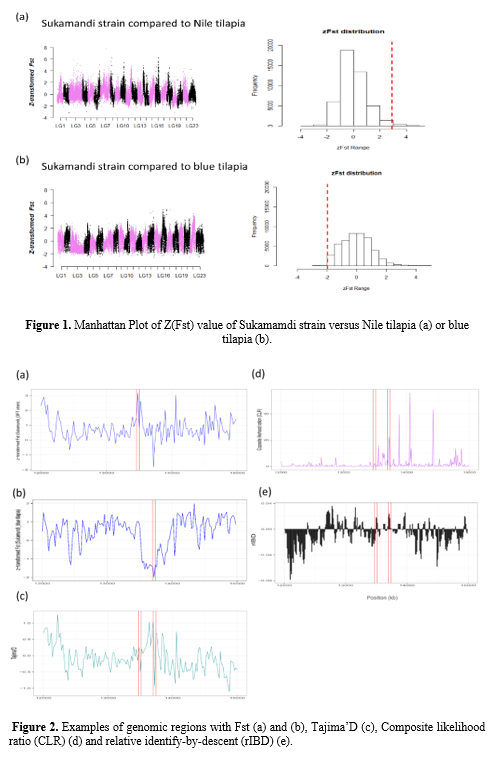
We compared the genome of this Sukamandi strain to that of Nile tilapia (Orechromis niloticus) and blue tilapia (Orechromis aureus ), the latter a putative donor of the salinity tolerance. Our results indicate that the Sukamandi strain is genetically much more similar to Nile tilapia (Orechromis niloticus) (Fst=0.042) than to blue tilapia (Orechromis aureus) (Fst =0.386). By two pairwise comparisons (as shown in Figure 1) , 33 salinity adaptive genes involved in MAPK3 activity, potassium ion homeostasis, ATPase activity (coupled to transmembrane movement of ions), calcium ion binding pathway were identified . C ombining genome-scale scanning for selection and introgression, revealed that salinity tolerance related genes, such as slc25a24 and cdh1 were under selection (as shown in Figure 2).
Conclusion
Salinity adaptive genes disclosed by genetic differentiation comparison with blue and Nile tilapia strain were partially under selection. The genome of Sukamandi strain appears to be overwhelmingly of Nile tilapia origin but introgression from blue tilapia may have conferred some of the observed salinity tolerance . We provide the genetic basis for breeding resilient tilapia -Sukamandi strain.