EFFLUENT-GROWN ULVA AS A FUNCTIONAL INGREDIENT FOR FARMED ABALONE: IMPACTS ON GROWTH, PHYSIOLOGY AND MICROBIOME
Introduction
Integrated Multi- Trophic Aquaculture (IMTA) represents a sustainable production method that can reduce the environmental impacts of aquaculture , facilitate diversification and increase production. S everal large-scale commercial abalone farm s in South Africa practice IMTA, growing Haliotis midae in land-based raceway tanks , interconnected with Ulva in adjacent paddle raceways using abalone effluent. Ulva serves as a biofilter, allowing water from the Ulva systems to be re-circulated back to abalone tanks, and Ulva is used as supplementary feed for abalone . Dietary Ulva supplementation has conveyed benefits to a variety of cultured animals , enhancing feed consumption, growth, product quality and health , with some of the improvements to abalone culture believed to be linked to the carbohydrate fraction of Ulva (Naidoo et al., 2006; Mulvaney et al., 2013; Kemp et al. 2015; Bansemer et al., 2016). The aim was to investigate the effects of (1) Ulva as a partial or complete replacement of formulated feed, and (2) the inclusion of Ulva, or specific components of Ulva, on feed consumption, growth, physiology and the gut microbiome of c ultured H. midae to gain a better understanding of the functional potential of Ulva as an aquafeed ingredient.
Materials and Methods
To test the effects of Ulva as a partial or complete replacement of a local formulated feed Abfeed®S34 (diet AB ) on total feed consumption, abalone (67.30±5.49g; n=10 per basket) were fed for 28 days with AB at a rate of 1.27% BW.day-1 or with graded levels of AB (75, 50, 25 & 0 % of AB); with the balance of the feed constituting of fresh IMTA grown Ulva (FU). A separate one-year on farm growth trial was conducted under farm conditions to assess the extent to which AB can be replaced by FU. Abalone (51.10±3.09g; n=200 per basket ) were fed AB at a rate of 0.27% BW.day-1 or graded levels of AB (100, 80, 70, 60 & 40% of AB ) supplemented with Ulva ( 0.21% BW.day-1, dry weight approximation). All treatments were offered in triplicate and growth and condition were assessed on days 112, 223 and 366.
A controlled laboratory trial was conducted t o test the effects of specific components of Ulva on abalone ( 20-30g) growth, physiology and gut microbiome. I sonitrogenous diets consisting of dried Ulva (10% w/w; AB10U ), Ulvan (1% w/w; AB1U ), and glucuronic acid (0.1% w/w; AB0.1U) were formulated. Feed conversion ratio (FCR), specific growth rate (SGR), tissue glycogen, and gut microbiome was assessed around day 0, 105 and 215 and compared to abalone fed diets AB, FU, or a combination of the latter (ABFU). The bacterial microbiome was characterised by sequencing the V3-4 hyper-varia ble region of the 16S rRNA gene. NGS was performed on an Illumina MiSeq sequencing platform, sequence data assessed using QIIME2 (Boylen et al., 2019), reads mapped against the SILVA 16S rRNA database (Quast et al., 2013) and summarized taxonomic abundance at different hierarchical levels were assessed.
Results and Discussion
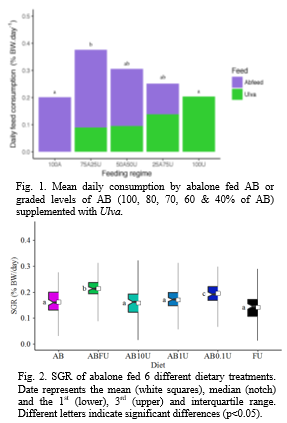
Incorporation of small amounts of fresh IMTA grown Ulva (25% ; diet 75A25U) were shown to significantly improve total feed consumption by ca. 90% , compared with abalone fed diets AB and FU (
1). N o significant differences in SGR , CF and tissue glycogen content were recorded between treatments in the 1 year growth trial , suggesting as much as 60% of a formulated feed can be substituted with FU without negatively affecting growth and condition of abalone.
The controlled laboratory trial, showed that abalone maintained on AB FU and AB0.1U had the highest SGR, significantly higher than abalone fed AB, AB10U, and FU (Fig. 2). Ab alone maintained on FU alone grew at a rate not statistically different to abalone maintained on AB , suggesting farmed abalone can be maintained on a diet consisting only of FU during the grow-out phase of production . Abalone fed diets supplemented with 0.1% glucuronic acid not only had improved SGR, but significantly lower FCR than abalone fed AB ; suggesting glucuronic acid may be one of the components within Ulva contributing towards growth. NMDS analysis of microbiome data revealed that abalone fed FU diets, and its components, produced significant associations in their intestinal bacterial communities, suggesting specific bacterial species are selected for and are associated with the digestive tract of abalone fed components of Ulva. Using DESeq2, s everal differentially abundant OTUs were identified across dietary treatments , with various bacterial genera, including members of the genus Vibrio found to be less abundant in the gut of abalone fed FU supplemented diets compared to AB alone. This study has demonstrated that dietary supplementation with IMTA- grown Ulva can reduce an abalone producer’s reliance on fishmeal-based dry formulated feeds and have several other health benefits.
References
Bansemer, M.S., Qin, J.G., Harris, J.O., Duong, D.N., Hai, T., Howarth, G.S. , Stone, D.A.J. 2016. Growth and feed utilisation of greenlip abalone ( Haliotis laevigata ) fed nutrient enriched macroalgae. Aquaculture. 452 , 62–68.
Bolyen, E., Rideout, J.R., Dillon, M.R., Bokulich, N.A., Abnet, C.C., Al-Ghalith, G.A., Alexander, H., et al. 2019. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology 37 , 852–857.
Kemp, J.O.G., Britz, P.J., Agüero , P.H.T. 2015. The effect of macroalgal, formulated and combination diets on growth, survival and feed utilisation in the red abalone Haliotis rufescens . Aquaculture. 448(C) , 306–314.
Mulvaney , W.J., Winberg, P.C., Adams, L. 2013. Comparison of macroalgal (Ulva and Grateloupia spp.) and formulated terrestrial feed on the growth and condition of juvenile abalone. Journal of Applied Phycology. 25(3) , 815–824.
Naidoo, K., Maneveldt, G., Ruck, K., Bolton, J.J. 2006. A c omparison of v arious seaweed-b ased d iets and f ormulated f eed on g rowth r ate of a balone in a land-b ased a quaculture s ystem. Journal of Applied Phycology. 18(3–5) , 437–443.
Quast, C. , Pruesse, E., Yilmaz, P. , Gerken, J. , Schweer, T. , Yarza, P. , Peplies, J. , Glöckner, F.O. 2013. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research 41(D1), D590–D596.