TIME-COURSE RNA-SEQ ANALYSIS OF BRAIN AND HEAD KIDNEY RESPONSE TO NERVOUS NECROSIS VIRUS INFECTION IN THE EUROPEAN SEA BASS
Introduction
Nervous Necrosis Virus (NNV) is one of the major vira l pathogens in aquaculture, affecting a wide range of fish species and causing high mortality rates . T he European sea bass (Dicentrarchus labrax) aquaculture production has long been affected , with significant economic losses, especially in the Mediterranean region. Selective breeding for increased resistance to NNV might help, together with other approaches (e.g. vaccination), to prevent such devastating disease. One of the major goals of the EU-funded project AQUAFAANG is the functional genomic characterization of important traits, which in turn might improve the accuracy of breeding value prediction. Here we present a detailed time-course RNA analysis on five time points and two tissues in juvenile sea bass, which is part of an integrated study to understand the genetic basis of response to NNV in this species.
Materials and methods
The experimental fish were generated at a commercial hatche ry Valle Ca’ Zuliani . Unselected juvenile fish (weight 6-20gr) were transferred to the IZSVe facility. After acclimation and quarantine, half of the animals were infected by intramuscular inoculum of 0,1ml RGNNV strain 283.2009 batch 5/19 at titre 106.30 TCiD50/ml. The remaining fish were injected with DMEM. All fish were anesthetised before injection with MS222. Four fish per time point per group (NNV-infected, mock-infected ) were sampled, euthanized, and brain and head kidney samples were dissected and stored for RNA analysis. Tissue sampling was carried out at 6h, 12h, 24h, 48h, and 72h post-inoculum. T otal RNA was extracted from all samples , quality assess using Agilent 2100 Bioanalyzer , and only RNAs with RIN >7 were processed. cDNA libraries were constructed using QuantSeq 3’ mRNA-Seq Library Prep Kit FWD for Illumina and sequenced on a HighSeq 4000 with a single 1 x 100 bp setup. High-quality trimmed reads were mapped on the European sea bass genome, (seaba ss_V1.0 GCA_000689215.1 in Ensembl). Normalisation was performed using the function RUVs (with in the package RUVSeq/v1.18. N ormalised counts were imported in edgeR /v3.26.8 to perform differential expression analysis in pairwise comparisons (time-matched mock-infected vs NNV-infected). Genes with an FDR adjusted p-value < 0.05 and FC>±1.5 were considered as differentially expressed (DEGs). Putative zebrafish orthologues to sea bass genes were obtained from Ensembl . Annotations from zebrafish orthologues were used for functional enrichment analysis in DAVID.
Results
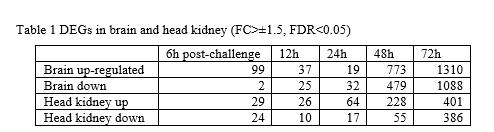
Transcriptomic response to NNV in both tissues was significant already at 6h post-inoculum and became massive at 48h and 72h. The number of up- and down-regulated genes in each tissue is reported in Table 1.
While the transcriptome response to NNV was quite complex, functional enrichment analysis showed a few general trends. Gene Ontology (GO) terms and KEGG pathways related to immune response (e.g. GO:0051607~defense response to virus , dre04622:RIG-I-like receptor signaling pathway ) and proteolysis/proteasome (e.g. GO:0006508~proteolysis , dre03050:Proteasome ) were significantly enriched (FDR<0.05) in up-regulated genes at 24h, 48h, 72h in both tissues. Down-regulated genes in the brain were enriched in GO terms and KEGG pathways related to vision (e.g. GO:0007601~visual perception , dre04744:Phototransduction ) as early as 12h after infection. In the head kidney, genes involved in glycolysis were significantly under-expressed in infected fish at 6h, while protein translation related genes (e.g. ribosomal genes) later on. The most relevant result was the significant involvement of different families of heat shock proteins (HSP40, HSP70, HSP90) in the host response to virus with up to 22 HSPs being differentially expressed. Of particular interest are HSP90ab1, HSP90b1, and HSP90aa1 , which showed a complex, tissue-specific pattern being up- and down-regulated at different time-points.
Discussion
Detailed time-course analysis of the brain and head kidney transcriptome revealed a vast, complex response, providing a large set of functi onally relevant genes to inform genomic prediction of NNV-resistance in the European sea bass. Of particular interest are HSP90s as HSPs are known to have a key role in host-virus interactions (Wan et al. 2020) and HSP90ab1 has been reported to be essential for RGNNV entry into target cells in marine medaka (Zhang et al. 2020).
References
Wan Q, Song D, Li H, He ML. Stress proteins: the biological functions in virus infection, present and challenges for target-based antiviral drug development. Signal Transduct Target Ther. 2020 Jul 13;5(1):125.
Zhang W, Jia K, Jia P, Xiang Y, Lu X, Liu W, Yi M. Marine medaka heat shock protein 90ab1 is a receptor for red-spotted grouper nervous necrosis virus and promotes virus internalization through clathrin -mediated endocytosis. PLoS Pathog. 2020 Jul 8;16(7):e1008668.