DIFFERENCES IN THE CIRCULAR RNA TRANSCRIPTOME BETWEEN WILD AND CAPTIVE-BRED NILE TILAPIA
Introduction
Domestication of organisms started as early as in the Neolithic period (c.14 000 years ago) . Historically, f ish domestication began almost 1000 years later than man y other species, such as cattle, dogs, pigs, and horses . Domestication is a complex and long process in which organisms are subject to human control in many aspects, including feeding and breeding, thus changing their phenotype compare to their wild ancestors. Accumulating shreds of evidence suggest that domestication plays crucial roles in modulating behaviour , size, colouration, morphology, and physiology in teleosts (Balon et al. 2004). These modifications are often linked to changes in genetic structure through mutations and allele fixation . Investigations into more recently domesticated species reveal that phenotypic change is relatively fast, and expression of hundreds of gene can be altered in a single generation. Epigenetic modifications , potentially heritable modifications of the chemical structure of the genome without affecting the nucleotide sequence, are important in the shaping phenotypic differences during the domestication process. The role of m ethylation, hydroxymethylation in domestication of fish has been described in artificial selection, environmental adaptation, and genome evolution (Podgorniak et al. 2019; Konstantinidis et al. 2020). In addition, non- coding RNAs can also influence phenotypic diversity and fitness by regulating transcription mechanisms and give rise to the emergence of domestication traits. Circular RNAs (circRNAs) are a class of endogenous non-codin g RNAs that have attracted interest in transcriptional regulation. They act as miRNA sponges and play a regulatory role in normal physiology as well as in pathological conditions. Despite the importance of circRNAs in several biological processes , their role in fish domestication remains largely unknown. In the present study , we compared wild and first-generation Nile tilapia (Oreochromis niloticus) reared in captivity to determine the potential role of circRNAs in domestication.
Materials and methods
The RNA-seq dataset for the wild and domesticated group (twelve samples) were obtained from ribosomal RNA depleted RNA-seq libraries published by Konstantinidis et al. (2021) under accession number GSE135811 in Gene Expression Omnibus. After filtering low-quality reads and adapters, sequences were mapped into the Nile tilapia reference genome (ASM185804v2). CircRNA prediction was performed for each RNA dataset with the circRNA in silico prediction tools CIRI2. Differential expression of circRNAs between the groups was assessed using circMeta (Chen L. et al., 2020) and the host genes of circRNAs were predicted using circParser (Artem et al., 2020).
Results
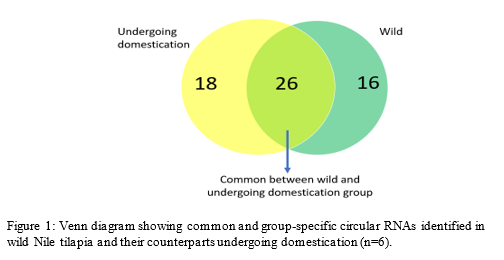
A total of 7,862 circRNAs were identified from a muscle RNA-seq dataset (n=6) using the CIRI2 algorithm. Among them, 26 circRNAs were present in both wild fish and their progeny undergoing domestication (Fig. 1). Further analysis revealed 39 differentially expressed (DE) circRNAs . Several DE circRNAs originate from host genes known to be involved in domestication, such as MHC class I and glutamate receptor-interacting protein 2. Furthermore, a number of DE circRNAs orthologues were found to be conserved between many fish species. Taken together, our findings reveal a potential role for circRNAs in fish domestication .
Acknowledgements
This study has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme [grant agreement no 683210] and from the Research Council of Norway under the Toppforsk programme [grant agreement no 250548/F20].
References
Balon, E.K., 2004. About the oldest domesticates among fishes. Journal of fish Biology, 65, pp.1-27.
Chen, L., Wang, F., Bruggeman, E.C., Li, C. and Yao, B. (2020). circMeta: a unified computational framework for genomic feature annotation and differential expression analysis of circular RNAs. Bioinformatics, 36(2), pp.539-545.
Konstantinidis, I., Sætrom, P., Mjelle, R., Nedoluzhko, A.V., Robledo, D. and Fernandes, J.M. (2020). Major gene expression changes and epigenetic remodelling in Nile tilapia muscle after just one generation of domestication. Epigenetics, 15(10), pp.1052-1067.
Nedoluzhko, A., Sharko, F., Rbbani, M.G., Teslyuk, A., Konstantinidis, I. and Fernandes, J.M. (2020). CircParser: a novel streamlined pipeline for circular RNA structure and host gene prediction in non-model organisms. PeerJ, 8, p.e8757.
Podgorniak, T., Brockmann, S., Konstantinidis, I. and Fernandes, J.M., 2019. Differences in the fast muscle methylome provide insight into sex-specific epigenetic regulation of growth in Nile tilapia during early stages of domestication. Epigenetics, 14(8), pp.818-836.