UNRAVELLING THE IMPACT OF INTESTINAL MICROBIOTA ON HOST TRANSCRIPTOMICS IN GILTHEAD SEA BREAM JUVENILES FED NON-CONVENTIONAL FEED INGREDIENTS
Introduction
In a sustainable aquafeed scenario, processed animal proteins (PAP), insect , microbial and algae-products appear a s suitable replacers of fish meal (FM) in practical fish diets ( Glencross et al., 2020; Basto et al., 2021) . However, such new and emerging ingredients have been tested one-by-one rather than using different formulation combinations . In this regard, a previous study in gilthead sea bream (Sparus aurata) pointed out that graded levels of sustainable PAP- and NoPAP - based feed formulations are able to support optimal growth performance when nutrient requirements are met (Fernandes et al., 2021). However, PAP -based feed formulations caused a slight impairment of feed conversion ratio (FCR) associated with a down-regulation of the hepatic insulin-growth factor-I and the up-regulat ion in head-kidney of pro-inflammatory cytokines, chemokines and T-cell markers. Here, we focused on gut health indicators to assess the main effects of PAP and NoPAP- based diets on the mucosal adherent microbiota of the anterior intestine (AI) and its impact on changes in the host transcriptomic profile of selected markers of intestinal function and integrity.
Methods
Quadruplicate groups of gilthead seabream were fed ad libitum daily with three different diets in a 77 days feeding trial . The control diet (CRTL) followed a commercial type formulation. In both NoPAP and PAP diets , insect meal, fish by-products, microbial and yeast biomasses were used as FM and vegetable protein replacers. T he PAP diet also comprised several PAP ingredients such as poultry meal, feather meal hydrolysate, and porcine blood meal. T he NoPAP diet included Spirullina and Chlorella meal as additional protein sources. At the end of the feeding trial , RNA from the AI was collected and run through a PCR-array to profile the expression of a panel of 44 genes, including markers of epithelial integrity, interleukins and immunoglobulins , among others. DNA from the adherent bacteria of the AI was collected and t he V3-V4 region of the 16S rRNA of each sample was amplified and sequenced by Illumina MiSeq. T axonomic assignment was performed against the RDP database. Alpha diversity was calculated using Phyloseq , and beta diversity using partial least-squares discriminant analysis (PLS-DA) models . Metagenome prediction and pathway analysis were performed using Piphillin. Differentially expressed (DE) genes and discriminant OTUs were correlated using the corrplot R package.
Results
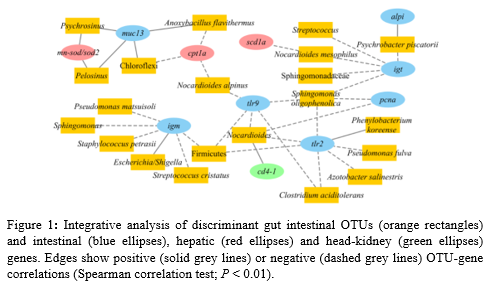
The gene expression analysis revealed 13 DE genes (out of 44) (P < 0.1) in response to the experimental diets. M arkers of epithelial integrity (pcna ) and pro-inflammatory genes (alpi , il8 and igm) were significantly up-regulated in fish fed PAP diet. Fish fed NoPAP diet presented an up- regulation of the anti-inflammatory cytokine il10. In the microbiota analysis, Illumina sequencing reads were assigned to 2,180 OTUs and a significantly lower richness and alpha diversity were found in the NoPAP group in comparison to CTRL fish. Detailed differences in microbiota composition were analysed with a statistically validated PLS-DA which clearly separated CTRL fish from fish fed PAP and NoPAP diets , with 135 OTUs mainly driving this separation (VIP ≥ 1). Inferred metagenome results showed a differential regulation of 3 4 pathways. Both NoPAP and PAP groups showed, among others, the up-regulation of routes tailoring immune response and inflammation ( C-type lectin receptor, VEGF, TNF and NFκ-β signalling pathways ), with a lower degree of activation in fish fed NoPAP diet. Fish fed PAP and NoPAP diets also displayed an exclusive type of response at this level, with the differential regulation of 14 and 10 inferred pathways, respectively. Correlation tests disclosed a remarkable amount of DE intestinal genes (12) involved in 38 significant (P < 0.01) correlations with 29 of the discriminant OTUs (Fig. 1). This highlights the relation between host gene expression and changes in the gut microbiota. In addition , 11 out of the 29 correlated OTUs were also associated with changes in the relative expression of liver and head-kidney genes, retrieved from previous studies performed in the same fish.
Conclusions
Even though growth performance was not affected by the experimental diets, these results disclosed a pro-inflammatory response to PAP -based formulation in terms of gene expression and intestinal microbiota . This inflammatory condition seems to be ameliorating using NoPAP -based feed formulation , which becomes especially interesting as an alternative eco-efficient feed for seabream . At the same time, a remarkable correlation between changes in gut microbial population and gene expression was unravelled at local and systemic levels . This highlights the potential action of the gut microbiome as a “second genome”, probably being involved in the regulation of the transcriptomic response of this marine farmed fish when fed innovative diet formulations based on increased circularity and resource utilization principles.
References
Basto et al., (2021). Frontiers in Physiology, 12, 659567 . doi : 10.3389/fphys.2021.659567
Fernandes et al., (2021 ). Presented at: EAS Aquaculture Europe 2020 Virtual, April 2021.
Glencross et al., (2020). Fishes 5, 1–39. doi: 10.3390/fishes5030022
Funding: GAIN (EU-H2020 #773330) ; AQUAEXCEL2020 (EU-H2020 #652831, T NA project AE150004); Bream-AquaINTECH (RTI2018–094128-B-I00 ); RYC2018-024049-I/AEI/10.13039/501100011033.